What happens when your healthcare data is used to train AI models?
If a single patient’s health data is included in development of an AI algorithm, how will this impact their care if they were to encounter that same algorithm in the future?
In recent years, increased focus has been given to improving the representativeness of training and validation datasets for healthcare algorithms. This means having more examples of all patient demographics, such as different ages, genders and ethnicities included during development, leading to better performance for everyone when a tool is deployed into the clinic.
This push for more diverse and representative data has sprung from a large body of research documenting the potential for biases against certain groups to lurk beneath what appears to be a high performing AI tool. These biases can be difficult to spot and can affect only certain subgroups of the population, a phenomenon known as hidden stratification.
There are several potential solutions to help models ‘generalise’ to all patient subgroups, but the best is to improve the numbers of minority groups in development datasets. The idea being that if your health data is included in algorithm development, then the resulting algorithm will perform better on patients like you in future. This is logical and an obvious example would be the inclusion of patients with darker skin tones in development of AI tools to detect skin cancer.
Whilst we might agree that inclusion of a patient from a given demographic group will improve performance for similar patients in the future, what is less certain is how it might impact future care for that very same patient. If a single patient’s health data is included in development of an AI algorithm, how will this impact their care if they were to encounter that same algorithm in the future?
Example scenario
This concept is a little abstract so is perhaps best explained with a example scenario…
2018 — our patient falls off their bike and has a suspected fractured rib. They are sent by their GP to get a chest X-ray which is reported by a radiologist and thankfully shows no signs of a fracture. They are sent home and recover quickly.
2020 — the hospital shares an anonymised set of 100,000 chest X-rays with an AI company looking to develop an algorithm to help speed up reporting by radiologists. Our patient’s X-ray from 2018 is included. The company labels the scan as ‘normal’ and includes it in model training. The eventual model is deployed to the hospital to assist radiologists, with good results.
2023 — our patient has developed a persistent cough and is again sent for a chest X-ray by their GP. This time, the X-ray shows the early signs of lung cancer. The AI algorithm has learnt from training data to recognise the small nodule and flag it to the radiologist, however it has also seen this patient before, it recognises the bone structure and lung features and was told during training that they are ‘normal’.
Will the AI tool successfully identify the new and deadly feature in the scan, or simply remember the label that was assigned to the patient’s earlier scan?

Why this matters:
This might all sound very theoretical, but here are three reasons why I think this could be a larger problem than it initially seems:
1. Use cases are getting more specific:
As AI in healthcare matures, use-cases become more specific, narrowing the group of patients that can be used for training and that might benefit from the tool when deployed. In particular, if patients tend to live with the disease for a long time, they have an increased likelihood of meeting an algorithm to which their data has contributed to the training of.
2. Shorter development times and model retraining:
It is taking less and less time to develop, test, regulate and deploy a healthcare algorithm. Whilst in the past health data collection might have had clinical impact on the timescales of decades, now this can be expected in months to years, making it more likely those same patients will still be battling a disease by the time an AI algorithm is deployed.
As models are deployed, they will also be closely monitored. Difficult cases are collected, labelled and added to datasets for retraining. A new version of the algorithm can then be deployed in a matter of months. This could be particularly relevant for repeat procedures, such as breast cancer screening, with women being screened every 3 years in the UK. It is not unlikely that in the time between mammograms, a patient’s scan could be collected, labelled and added to training — potentially influencing the interpretation of her next scan.
3. Foundation models:
Until now, developers have largely relied on models that are pre-trained using general image datasets. However, self-supervised learning techniques are making task-specific pre-trained networks more popular. This means developers can achieve higher performance because their models already know what a chest X-ray is, they just need to point out what to look for. This is great and could have huge implications for transferability of algorithms between clinical settings, but it also means larger patient cohorts will be included in training of algorithms. Concerns have already been raised about the potential for these foundation models to encode racial and social bias. It is uncertain how much of a footprint individual inclusion in foundation model training might leave and if this will be carried forward to fine-tuned tasks.
Experiment Setup:
To investigate this question, I will train a deep learning model to classify chest X-rays using the public CXR14 dataset. This consists of 114,000 X-rays from 30,000 patients, all labelled with 14 common findings. Many patients have several scans which can be ordered chronologically by the follow-up number provided.
Our model is trained using a subset of 10,000 scans (one per patient), a test dataset was constructed with 2,000 scans (one per patient) and validation performed on a held-out set of 4,000 scans. This validation set is built to answer our specific question and therefore contains three main patient groups:
● ‘Unseen’ Patients (2,000 scans) — none of these patients were included in the 10,000 used for training.
● ‘Seen and Unchanged’ Patients (1,000 scans) — earlier scans from these patients were included in training dataset but the label assigned to the scans is the same (unchanged).
● ‘Seen and Changed’ Patients (1,000 scans) — earlier scans from these patients were included in training dataset and the label assigned to the scans is different (changed).
Models were trained with a standard setup using the PyTorch Lightning library. A ResNet-18 model is trained as the baseline and ResNet-34 trained to investigate the impact of greater model capacity.
Model performance is compared between the 3 validation dataset groups with classification of all positive labels (i.e. abnormality detection) being considered for calculation of AUC values.
Results:
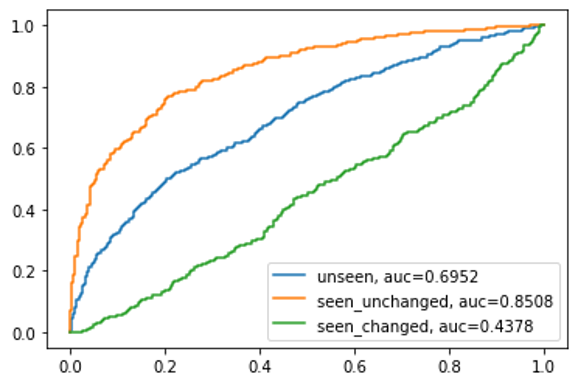
The ROC curve and AUC values for the three patient validation groups for the ResNet-18 model can be seen below (see here for how to interpret a ROC curve) . The AUC recorded for unseen patients across all labels was 0.69, this is in line with the recorded performance on the test dataset of 0.706 (not shown on graphic). So far, so expected.

Where things become less expected is if we look at the performance of the model on the patients who were ‘Seen’ — i.e. they had a prior scan included in model training. For the group of patients who were unchanged, meaning the second scan had the same label as the first, our model performs significantly better than on unseen patients, with an AUC of 0.8256.
When considering ‘seen and changed’ patients, whose second scan had a different label to the first, performance is significantly worse. In fact, an AUC value of 0.436 is actually worse than totally random classification (which would give a 0.5 value). This suggests the model could be predisposed to simply repeat the previous label for a patient and this is competing with clinically relevant features in the image.
Score Distributions:
Delving deeper, we can visualise the overall score distributions amongst normal and abnormal scans. If our model is performing well, we should see a clear separation between the classes. When visualising this for unseen patients, there is a clear peak of normal scans (blue) with low overall abnormality scores. An even clearer separation can be seen for ‘seen and unchanged’ scans, with the model better able to identify positive cases. However, for those scans from ‘seen and changed’ patients, there is virtually no separation between classes, furthering the suggestion that this model is no better than random for this group.

Does model size change things?
One theory to explain this behaviour might be that our model does not have enough capacity to learn clinically relevant features and so relies on overfitting to more general patient features. To test this, I trained a ResNet-34 to complete the same task, with more than twice as many parameters to train.
However, the ROC curve and AUC values below show the same pattern between the three patient groups. In fact, the disparities between changed and unchanged patients is even larger than with the ResNet-18 baseline.
It is not entirely clear why this is the case. It could be that our model is actually overfitting and the larger model does this to a greater extent. Or, that the increased model capacity offers more potential for individuals to be memorised and therefore training labels simply repeated back.
Conclusions
Any conclusions to be taken away from this are naturally limited by the fact that this is a small experiment with many other factors to be considered. However, we do observe consistent behaviour of models appearing to memorise training labels for individual patients, meaning changes between prior and current scans could be ignored or over-ruled.
If this behaviour is common across diagnostic AI solutions it is likely missed by virtually all clinical AI validations, as training and validation datasets are routinely split by patient index. Meaning the scenario of a prior scan being present in training cannot occur in algorithm validations required by regulators.
As outlined, this scenario has the potential to increase in frequency as AI is more widely deployed and utilised within healthcare and could risk significant harm if newly developed signs of disease are overlooked by otherwise trusted AI models.